Publications
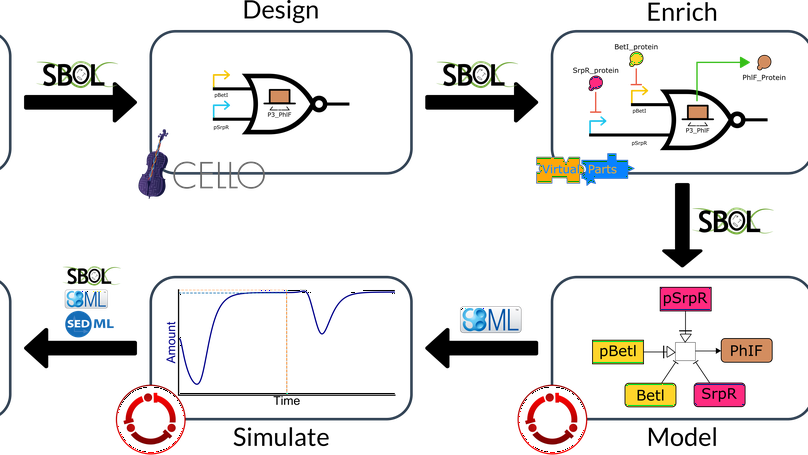
Multiple input changes can cause unwanted switching variations, or glitches, in the output of genetic combinational circuits. These glitches can have drastic effects if the output of the circuit causes irreversible changes within or with other cells such as a cascade of responses, apoptosis, or the release of a pharmaceutical in an off-target tissue. Therefore, avoiding unwanted variation of a circuit’s output can be crucial for the safe operation of a genetic circuit. This paper investigates what causes unwanted switching variations in combinational genetic circuits using hazard analysis and a new dynamic model generator. The analysis is done in previously built and modeled genetic circuits with known glitching behavior. The dynamic models generated not only predict the same steady-states as previous models but can also predict the unwanted switching variations that have been observed experimentally. Multiple input changes may cause glitches due to propagation delays within the circuit. Modifying the circuit’s layout to alter these delays may change the likelihood of certain glitches, but it cannot eliminate the possibility that the glitch may occur. In other words, function hazards cannot be eliminated. Instead, they must be avoided by restricting the allowed input changes to the system. Logic hazards, on the other hand, can be avoided using hazard-free logic synthesis. This paper demonstrates this by showing how a circuit designed using a popular genetic design automation tool can be redesigned to eliminate logic hazards.